The A1 allele of the TaqI-A polymorphism, located 9.4 Kb downstream from the coding region of the DRD2 gene, has been associated with a variety of phenotypes/endophenotypes, among them, severe alcoholism and psychopathic traits in alcohol-dependent patients. Reference Noble1,Reference Hoenicka and Aragues2 However, a functional polymorphism of the DRD2 gene (C957T) has been proposed as being responsible for some of the phenotypic features which had been previously associated with the A1 allele, such as D2 dopamine receptor availability in the striatum. Reference Hirvonen, Laakso, Nagren, Rinne, Pohjalainene and Hietala3 The C957T DRD2 single nucleotide polymorphism (SNP) affects messenger RNA stability and protein synthesis in vitro. Reference Duan, Wainwright, Comeron, Saitou, Sanders, Gelernter and Gejman4 Additionally, it is now known that the TaqI-A polymorphism is located in a nearby novel gene named Ankyrin Repeat and Kinase Domain Containing (ANKK1) where it causes a missense substitution. Reference Neville, Johnstone and Walton5 A question that remains to be answered is which other phenotypes previously associated with the A1 allele are better explained by the proximity to the C957T DRD2 polymorphism, and which are related to ANKK1 itself. Interestingly, the TaqI-A SNP of the ANKK1 gene, but not C957T of the DRD2 SNP, is associated with increased striatal activity of aromatic L-amino acid decarboxylase, the final enzyme in the biosynthesis of dopamine. Reference Laakso, Pohjalainen, Bergman, Kajander, Haaparante, Solin, Syälahti and Hietala6
From the clinical point of view, a major issue is the relationship of A1+ genotype with antisocial behaviours, Reference Hoenicka and Aragues2,Reference Ponce, Jimenez-Arriero, Rubio, Hoenicka, Ampuero, Ramon and Palomo7 given the importance of antisocial personality disorder in patients with alcoholism and other substance misuse disorders. Reference Regier, Farmer, Rae, Locke, Keith, Judd and Goodwin8,Reference Helzer and Pryzbeck9 It has been proposed that the concept of psychopathy, as defined by Hare's Psychopathy Checklist–Revised (PCL–R), Reference Hare10 would be helpful for the identification of a more homogeneous group among alcohol-dependent patients with behavioural disturbances. This concept, based in Cleckley's criteria, Reference Cleckley11 places more weight upon the psychological nuclear characteristics rather than non-adaptive behaviours.
Taking into account all this information, we have investigated the hypothesis that the risk for psychopathic traits in alcohol-dependent patients may be related to both TaqI-A ANKK1 SNP and C957T DRD2 SNP.
Methods
Participants
The study included 176 alcohol-dependent Spanish males who had consecutively requested detoxification at the Addictive Behaviour Unit of the ‘Doce de Octubre’ University Hospital. Inclusion criteria were (a) males aged 18–65 years and (b) alcohol dependence according to DSM–IV criteria. 12 Exclusion criteria included the presence at baseline evaluation of another axis I psychiatric disorder diagnosed according to DSM–IV Reference First, Spitzer, Gibbon and Williams13 (except nicotine addiction). In addition, a number of other more general factors were considered such as the absence of family relationships and family history of immigration. Both historical data and immunogenetic studies have previously shown that the Spanish population is ethnically homogeneous and has been stable for at least 500 years. Reference Arnaiz-Villena, Martinez-Laso and Alonso-Garcia14 Patients of other ethnic backgrounds were not included in this study. The general characteristics of the sample patients were as follows: mean age, 42.1 years (s.d.=10.7); mean age at misuse onset, 23.2 years (s.d.=7.0); mean age at dependence onset, 30.1 years (s.d.=9.1); mean grams of alcohol per intake, 224.2 g (s.d.=166.2); number of patients with physiological dependence, 145 (82.4%); number of patients with family history of alcoholism, 97 (55.1%); and number of patients with alcohol-related medical problems, 79 (44.8%).
A group of 150 healthy individuals were recruited as controls on whom a genetic study was also performed following a clinical interview to ensure non-dependence on alcohol or other drugs. All alcohol-dependent patients and controls were unrelated to individuals within their group.
The study was approved by an ethics committee and informed written consent was obtained from all participants.
Assessment of psychopathy and antisocial behaviour
In order to evaluate psychopathic traits in the patients, the PCL–R Reference Hare10 was used as described in Hoenicka et al. Reference Hoenicka, Ponce, Jiménez-Arriero, Ampuero, Rodriguez-Jiménez, Rubio, Aragüés, Ramos and Palomo15 We applied the version that had previously been standardised by Moltó et al Reference Moltó, Poy and Torrubia16 for its use in male prison inmates in Spain. Given that cut-off values have not been established for the PCL–R scale in populations of alcohol-dependent patients, this scale was not employed to make a diagnosis of psychopathy. The PCL–R was, therefore, used in a dimensional manner in order to evaluate the degree of association of genetic markers with the total score across the whole scale. The PCL–R scale sub-factors were also evaluated dimensionally. Reference Reardon, Lang and Patrick17
We used criteria from ICD–10 18 for the diagnosis of dissocial personality disorder as a categorical approach, since it has been proposed that the attributes measured by the PCL–R are similar in many respects to this disorder. Reference Hare, Clark and Grann19 The diagnosis of dissocial personality disorder was made by applying the International Personality Disorder Examination (IPDE; ICD–10 module). Reference Loranger, Janca and Sartorius20
Genotyping
High molecular weight genomic DNA was purified from peripheral blood leukocytes using standard methods. This DNA served as a template for subsequent analysis by polymerase chain reaction-based methods. Genotyping was performed by Taqman assays, designed to run on an ABI 7900HT machine with sequence detection system software. The sequence-specific primers for TaqI-A ANKK1 SNP, 5′-CTGCCTCGACCAGC-3′ and 5′-CTGCCTTGACCAGC-3′, were designed and used for the C (A2) and T (A1) allele respectively, in addition to a common reverse primer 5′-GCAACACAGCCATCCTCAAAG-3′. Ten samples for each genotype were confirmed by direct sequencing analysis. The resulting genotypes for the TaqI-A ANKK1 SNP were clustered according to the presence of at least one A1 allele (A1+ genotype: A1 allele homozygous and heterozygous; A1–genotype: homozygous for the A2 allele) as described by Blum et al. Reference Blum, Noble, Sheridan, Montgomery, Ritchie, Jagadeeswaran, Nogami, Briggs and Cohn21 European population estimates for A1+ genotype frequencies range from 20.8 to 43.4% (National Center of Biotechnology Information (NCBI), www.ncbi.nlm.nih.gov; identification number rs1800497).
The C957T DRD2 SNP was analysed as previously described by Hoenicka et al. Reference Hoenicka, Ponce, Jiménez-Arriero, Ampuero, Rodriguez-Jiménez, Rubio, Aragüés, Ramos and Palomo15 Briefly, the sequence-specific primers for Taqman assays 5′-CTGTCGGGAGTGCTG-3′ and 5′-CTGTCAGGAGTGCTG-3 were used for the C and T alleles respectively, and the common reverse primer 5′-GCCCATTCTTCTCTGGTTTGG-3′. The genotypes obtained were grouped assuming a recessive model for the C957 allele: homozygous individuals for the C allele v. heterozygous and homozygous individuals for the T allele. European population estimates for CC genotype frequency range from 12.5 to 17.2% (NCBI, identification number rs6277).
The Hardy–Weinberg equilibrium test with the Genetic Data Analysis software version 1.1 for Windows found no deviation from either controls (TaqI ANKK1 SNP, P=0.44; C957T DRD2 SNP, P=0.78) or patients (TaqI ANKK1 SNP, P=0.37; C957T DRD2 SNP, P=0.98).
Taking into account the size of our sample of patients and an α-value of 0.05, we calculated the power of the genotype study for the two SNPs using Power Sample Size version 2.1.31 software for Windows. Our study varies with OR=2.4 and a power of 86.6% for TaqI-A ANKK1 SNP, and OR=2.0 and a power of 80.6% for C957T DRD2 SNP.
Statistical analysis
To evaluate the non-random association of the two SNPs, pairwise linkage disequilibrium statistics D′ and correlation coefficient r 2 were calculated using Haploview software version 3.2 for Windows (Whitehead Institute for Biomedical Research, www.broad.mit.edu/mpg/haploview/index.php). D′ explains the difference in frequency between the observed and expected number of SNP pairs. Being scaled to D max, it spans the range −1 to 1, and r 2 is the squared correlation coefficient between the markers. We found a low degree of TaqI-A to C957T linkage disequilibrium (patients D′=0.576, r 2=0.13; controls D′=0.58, r 2=0.14).
Case–control association genetic study with alcoholism
To compare differences between alcohol-dependent patients and healthy controls with respect to the genotype for each SNP, a chi-squared test was employed.
Case–case association genetic study with psychopathic traits and dissocial personality disorder
In the sample of patients, in order to evaluate the effect of each genotype on the average score on the PCL–R scale, we performed analyses of covariance using age as a covariate, since it has been reported that PCL–R scores may vary with age. Reference Reardon, Lang and Patrick17
Chi-squared tests were employed to compare genotype frequencies between alcohol-dependent patients with and without a diagnosis of dissocial personality disorder. A stepwise logistic regression model was used to calculate the independent association of each gene with the presence of dissocial personality disorder. Odds ratios, 95% confidence intervals and Wald values were estimated.
Case–case interaction study
A post hoc regression analysis of the interaction between genotypes and the effect upon the presence of dissocial personality disorder was also carried out.
The total sample of alcohol-dependent patients was subdivided according to the number of risk genotypes in order to study the global effect on dissocial personality disorder and PCL–R scores. Again, the chi-squared test was used to compare the presence of dissocial personality disorder between these subgroups, and post hoc analyses of covariance to compare PCL–R scores.
In order to correct for multiple comparisons, a Bonferronicorrected α-level of P<0.025 (0.05/2) was used for the two independent variables examined in Table 1. Otherwise, an α-level of 0.05 was used throughout. These statistical calculations were performed using the SPSS Statistical Package, version 11.1D for Windows.
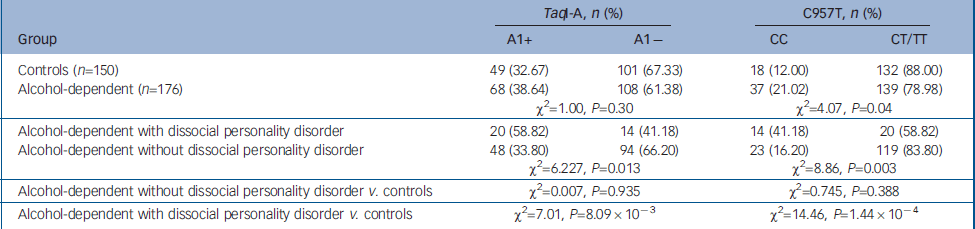
Table 1 Distribution of TaqI-A ANKK1 SNP and C957T DRD2 SNP genotypes in controls and alcohol-dependent patients
| TaqI-A, n (%) | C957T, n (%) | |||
|---|---|---|---|---|
| Group | A1+ | A1- | CC | CT/TT |
| Controls (n=150) | 49 (32.67) | 101 (67.33) | 18 (12.00) | 132 (88.00) |
| Alcohol-dependent (n=176) | 68 (38.64) | 108 (61.38) | 37 (21.02) | 139 (78.98) |
| χ2=1.00, P=0.30 | χ2=4.07, P=0.04 | |||
| Alcohol-dependent with dissocial personality disorder | 20 (58.82) | 14 (41.18) | 14 (41.18) | 20 (58.82) |
| Alcohol-dependent without dissocial personality disorder | 48 (33.80) | 94 (66.20) | 23 (16.20) | 119 (83.80) |
| χ2=6.227, P=0.013 | χ2=8.86, P=0.003 | |||
| Alcohol-dependent without dissocial personality disorder v. controls | χ2=0.007, P=0.935 | χ2=0.745, P=0.388 | ||
| Alcohol-dependent with dissocial personality disorder v. controls | χ2=7.01, P=8.09 × 10-3 | χ2=14.46, P=1.44 × 10-4 | ||
Results
Case–control association genetic study with alcoholism
Population estimates for CC and A1+ genotypes in our controls are comparable to those previously reported (see Methods). We then analysed the C957T DRD2 SNP and found a higher prevalence of the CC genotype in alcohol-dependent patients than in healthy controls. This difference did not reach statistical significance when multiple-comparison correction was applied. In the case of the TaqI-A ANKK1 SNP, we did not observe significant differences in the frequencies of the genotypes between alcohol-dependent patients and healthy controls (Table 2).
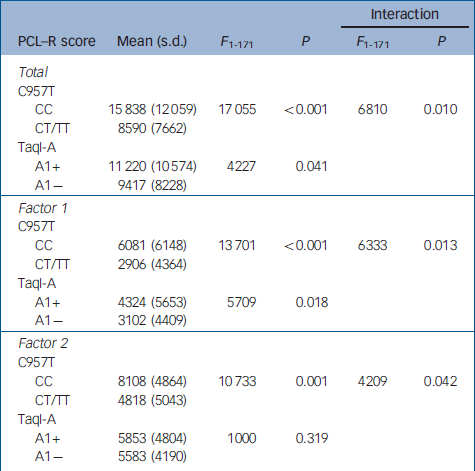
Table 2 Relationship between TaqI-A ANKK1 and C957T DRD2 SNP genotypes and PCL—R scores
| Interaction | |||||
|---|---|---|---|---|---|
| PCL-R score | Mean (s.d.) | F 1-171 | P | F 1-171 | P |
| Total | |||||
| C957T | |||||
| CC | 15 838 (12 059) | 17 055 | <0.001 | 6810 | 0.010 |
| CT/TT | 8590 (7662) | ||||
| TaqI-A | |||||
| A1+ | 11 220 (10 574) | 4227 | 0.041 | ||
| A1- | 9417 (8228) | ||||
| Factor 1 | |||||
| C957T | |||||
| CC | 6081 (6148) | 13 701 | <0.001 | 6333 | 0.013 |
| CT/TT | 2906 (4364) | ||||
| TaqI-A | |||||
| A1+ | 4324 (5653) | 5709 | 0.018 | ||
| A1- | 3102 (4409) | ||||
| Factor 2 | |||||
| C957T | |||||
| CC | 8108 (4864) | 10 733 | 0.001 | 4209 | 0.042 |
| CT/TT | 4818 (5043) | ||||
| TaqI-A | |||||
| A1+ | 5853 (4804) | 1000 | 0.319 | ||
| A1- | 5583 (4190) | ||||
Case–case association genetic study with dissocial personality disorder
When patients were separated according to whether they had comorbid dissocial personality disorder, the CC genotype for the C957T SNP was found to be overrepresented in the group of patients with dissocial personality disorder (OR=5.13), as well as the A1+ genotype for the TaqI-A SNP (OR=2.95). These differences were statistically significant (Table 1). Furthermore, no statistically significant differences were observed between healthy controls and alcohol-dependent patients without dissocial personality disorder.
A logistic regression analysis was then performed using the presence of dissocial personality disorder as a dependent variable, which confirmed that the A1+ genotype has a significant effect that cannot be explained by the presence of C957T (CC: OR=3.26, 95% CI 1.4–7.5, Wald=7.71, P=0.005; A1+: OR=2.51, 95% CI 1.1–5.5, Wald=5.27, P=0.022), and vice versa. The post hoc logistic regression analysis showed a significant pattern of interaction (OR=10.52, 95% CI 3.74–29.62, P<0.0001).
To gain more insight into the possible interaction between the DRD2 and ANKK1 genes, we studied the effect of carrying none, only one, or both risk genotypes identified in this study. The expression of dissocial personality disorder in individuals that were carriers of both risk genotypes (CC and A1+) was higher than that of patients who were carriers of none or only one risk genotype, either A1+ or CC (χ2=26.56, d.f.=3, P<0.001) (Fig. 1).
Case–case association genetic study with psychopathic traits
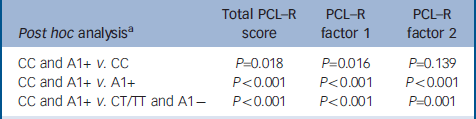
When the sample of patients was evaluated for genetic association with psychopathic traits, analyses of covariance revealed that the CC genotype of the C957T DRD2 SNP was associated with significantly higher scores on the PCL–R total and on both factor 1 and factor 2 of the scale. In contrast, the A1+ genotype of the TaqI-A ANKK1 SNP was associated with significantly higher scores only on factor 1. Moreover, the statistical analysis revealed a significant interaction effect between these two polymorphisms, and post hoc analyses (Scheffe) revealed that scores on the PCL–R and its factors were significantly higher only in the group of patients who were carriers of both CC and A1+ genotypes (Table 3), whereas there were not significant differences between the other three groups.
Table 3 Post hoc analyses between patients carrying only CC, only A1+, and CT/TT and A1- genotypes
| Post hoc analysisa | Total PCL—R score | PCL—R factor 1 | PCL—R factor 2 |
|---|---|---|---|
| CC and A1+ v. CC | P=0.018 | P=0.016 | P=0.139 |
| CC and A1+ v. A1+ | P<0.001 | P<0.001 | P<0.001 |
| CC and A1+ v. CT/TT and A1- | P<0.001 | P<0.001 | P=0.001 |
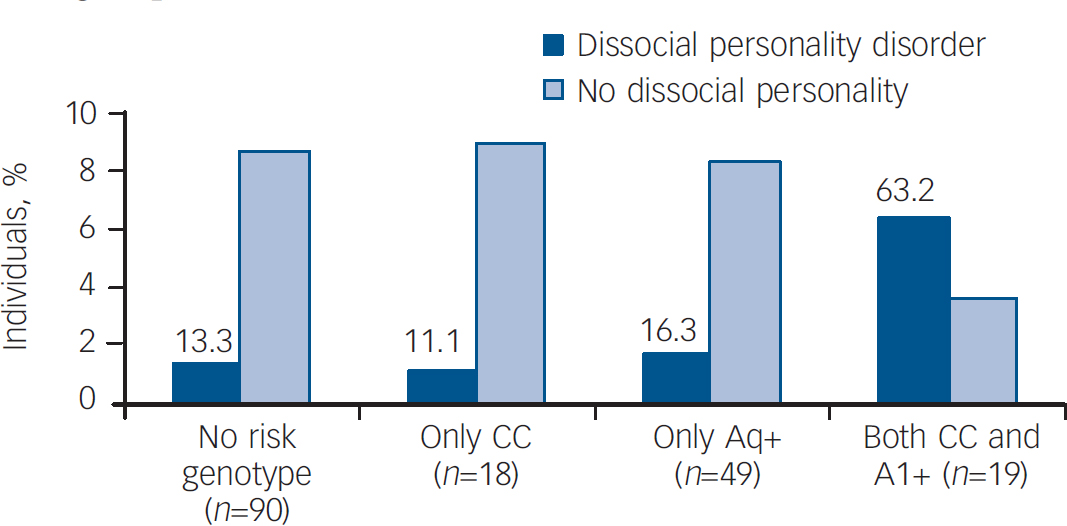
Fig. 1 Percentage of individuals who met criteria for dissocial personality disorder when none, only one or both risk genotypes were present.
The frequency of dissocial personality disorder is the highest in the group of carriers of both risk genotypes, reaching statistical significance. There are no significant differences between the other three groups.
Discussion
Main findings
We found CC and A1+ genotypes to be overrepresented only in alcohol-dependent patients with comorbid dissocial personality disorder. We also identified an association of both risk genotypes with high scores on PCL–R. Moreover, our results suggest that these genes epistatically influence the expression of dissocial personality disorder and psychopathic traits in our sample of alcohol-dependent patients.
This genetic interaction may reflect a functional relationship between the dopaminergic receptor D2 and the putative kinase ANKK1, given that it has been proposed that this protein could exert an effect on dopaminergic neurotransmission itself. Reference Laakso, Pohjalainen, Bergman, Kajander, Haaparante, Solin, Syälahti and Hietala6 Our results also illustrate the potential benefit of studying interactions between functional polymorphisms known to be involved in the same biological pathway, especially when studying multigenic disorders/traits. In a previous study, we have reported an additive relationship of the A1+ genotype, together with genetic variants for the CNR1 and FAAH genes, with psychopathic traits, using the PCL–R. Reference Hoenicka, Ponce, Jiménez-Arriero, Ampuero, Rodriguez-Jiménez, Rubio, Aragüés, Ramos and Palomo15 The new data presented here reveal that the effect of the A1+ genotype on the psychopathic phenotype depends on the interaction with the CC genotype for the DRD2 C957T SNP.
Implications
Our results regarding the association between the risk genotypes identified in this study and psychopathic traits in alcohol-dependent patients provides further support to previous family studies that have suggested that substance misuse disorders and antisocial behaviour could share, at least in part, common genetic vulnerability factors. Reference Cadore, Yates, Troughton, Woodworth and Stewart22 The psychopathy construct has demonstrated its validity since it correlates with variations in neurobiological function, Reference Deeley, Daly, Surguladze, Tunstall, Mezey, Beer, Ambikapathy, Robertson, Giampietro, Brammer, Clarke, Dowsett, Fahy, Phillips and Murphy23,Reference Birbaumer, Veit, Lotze, Erb, Hermann, Grodd and Flor24 and a recent heritability study has described that callous-unemotional traits are under strong genetic influence. Reference Viding, Blair, Moffitt and Plomin25 Psychopathy has been considered a condition based on differences in learning mechanisms Reference Blair26,Reference Newman, Kosson and Patterson27 suggesting the concourse of the mesolimbic dopaminergic system, which in turn has been repeatedly linked to addictive behaviours. Reference Di Chiara and Imperato28 Given the central role of dopamine in the reward system, the gene that encodes for the D2 dopamine receptor has been widely studied in relation to these disorders.
The change in the views regarding the implication of locus 11q23-24 with respect to phenotypic characteristics related to dopaminergic function has received support with the discovery of the ANKK1 gene, Reference Neville, Johnstone and Walton5 where the TaqI-A SNP is actually located. Recently, more extensive genotyping across DRD2 and ANKK1 confirms that the association with substance dependence and antisocial traits might be due to genetic variants in the ANKK1 gene. Reference Yang, Kranzler, Zhao, Gruen, Luo and Gelernter29,Reference Dick, Wang, Plunkett, Aliev, Hinrichs, Bertelsen, Budde, Goldstein, Kaplan, Edenberg, Nurnberger, Hesselbrock, Schuckit, Kuperman, Tischfield, Porjesz, Begleiter, Bierut and Goate30
The protein encoded by ANKK1 is one of a family of proteins involved in signal transduction pathways. Since TaqI-A, but not C957T, has been associated with differences in striatal activity of aromatic L-amino acid decarboxylase, Reference Laakso, Pohjalainen, Bergman, Kajander, Haaparante, Solin, Syälahti and Hietala6 ANKK1 could be related to dopamine pathways. The TaqI-A SNP causes an amino acid substitution within the 11th ankyrin repeat of the putative protein, which may affect substrate-binding specificity.
Combining these results with ours suggests that the fact that the ANKK1 and DRD2 genes are so close is not due to chance, instead that they could somehow interact in neurotransmitter pathways, affecting a person's vulnerability to both addiction and antisocial traits.
Study limitations
A limitation of our study derives from the way in which we applied the PCL–R. It must be noted that the PCL–R was not used for diagnosing psychopathy, but for evaluating specific traits related to antisocial behaviour. The PCL–R's reliability and specificity have been demonstrated in prison inmates in Spain. Reference Moltó, Poy and Torrubia16 Although few studies have been performed in substance-dependent patients, the PCL–R is generalisable across prison and substance-dependent samples, Reference McDermott, Alterman and Caccioca31 and recent studies have proposed that the psychopathy construct must be separated from criminality. Reference Cooke, Michie and Skeem32 On the other hand, the factorial structure of the PCL–R in different populations is still an open issue. Reference Cooke, Michie and Skeem32,Reference Johansson, Andershed, Kerr and Levander33
Given the relatively small size of our samples, our results should be considered as preliminary. To validate our results, a study of independent samples of alcohol-dependent patients, as well as populations of individuals free of addictive disorders, would be desirable, although there are obvious difficulties in recruiting samples of individuals with psychopathic personality disorders without confounding comorbidity. The analysis of families in which both alcoholism and antisocial disorders are present would also help to clarify the role of ANKK1 and DRD2 variants. In addition, we cannot rule out the existence of phenotypical differences in the A1A1 and the A1A2 genotype carriers. The size of our A1A1 patient sample is small, and hence could be deemed insufficient to detect a real contribution.
Despite these limitations, our study suggests that the expression of antisocial traits in alcohol-dependent patients relies, at least in part, on the existence of a potential interaction between ANKK1 and DRD2.
Finally, one cannot automatically infer biological interaction from statistical interaction, but if our findings are replicated, future research should also examine additional markers covering the two genes, as well as their intergenic linkage disequilibrium and haplotypes. Undoubtedly, functional studies of the ANKK1 gene, its putative peptide, and its relation with the D2 receptor, could open new avenues into the investigation of the mechanisms underlying the high comorbidity of antisocial personality disorder with alcoholism and other substance misuse disorders.
Acknowledgements
Supported by the Spanish Ministry of Health, Instituto de Salud Carlos III, CIBERSAM and by Fondo Investigaciones Sanitarias (FIS) grant no. 05/0731. We thank Carmen Aguirre for her invaluable collaboration in collecting blood samples and her empathic care of us and our patients.









eLetters
No eLetters have been published for this article.